You need to sign in or sign up before continuing.
Welcome to Git@WUR
Login
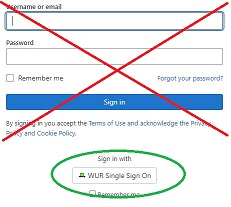
External users: use the username/password form.
WUR users: use the button 'WUR Single Sign On'.
Two-factor authentication (2FA)
Please enable two-factor authentication if not already enabled.
For general information visit the articles about GIT in the ‘WUR support portal'.
For technical information go to ‘Help files Git@WUR’.